-Search query
-Search result
Showing 1 - 50 of 77 items for (author: cohen & se)

EMDB-43931: 
CryoEM structure of activated CRAF/MEK/14-3-3 complex with NST-628
Method: single particle / : Quade B, Cohen SE, Huang X

EMDB-43932: 
Activated CRAF/MEK heterotetramer from focused refinement of CRAF/MEK/14-3-3 complex
Method: single particle / : Quade B, Cohen SE, Huang X

PDB-9axa: 
CryoEM structure of activated CRAF/MEK/14-3-3 complex with NST-628
Method: single particle / : Quade B, Cohen SE, Huang X

PDB-9axc: 
Activated CRAF/MEK heterotetramer from focused refinement of CRAF/MEK/14-3-3 complex
Method: single particle / : Quade B, Cohen SE, Huang X

EMDB-34806: 
SARS-CoV-2 Delta Spike in complex with FP-12A
Method: single particle / : Chen X, Wu YM

EMDB-34807: 
SARS-CoV-2 Delta Spike in complex with IS-9A
Method: single particle / : Mohapatra A, Wu YM

EMDB-34808: 
SARS-CoV-2 Omicron BA.1 Spike in complex with IY-2A
Method: single particle / : Chen X, Mohapatra A, Wu YM

PDB-8hhy: 
SARS-CoV-2 Delta Spike in complex with IS-9A
Method: single particle / : Mohapatra A, Wu YM

PDB-8hhz: 
SARS-CoV-2 Omicron BA.1 Spike in complex with IY-2A
Method: single particle / : Chen X, Mohapatra A, Wu YM

EMDB-12701: 
CryoEM structure of the transcription termination factor Rho from Mycrobacterium Tuberculosis
Method: single particle / : Martin K, Saridakis E, Vishwakarma R, Lai Kee Him J, Simon I, Cohen-Gonsaud M, Coste F, Margeat E, Boudvillain M, Bron P

PDB-7oqh: 
CryoEM structure of the transcription termination factor Rho from Mycobacterium tuberculosis
Method: single particle / : Saridakis E, Vishwakarma R, Lai Kee Him J, Martin K, Simon I, Cohen-Gonsaud M, Coste F, Bron P, Margeat E, Boudvillain M

EMDB-25209: 
Structure of human SARS-CoV-2 neutralizing antibody C1C-A3 Fab
Method: single particle / : Pan J, Abraham J, Yang P, Shankar S

EMDB-25210: 
Structure of human SARS-CoV-2 spike glycoprotein trimer bound by neutralizing antibody C1C-A3 Fab (variable region)
Method: single particle / : Pan J, Abraham J, Shankar S

PDB-7sn2: 
Structure of human SARS-CoV-2 neutralizing antibody C1C-A3 Fab
Method: single particle / : Pan J, Abraham J, Yang P, Shankar S

PDB-7sn3: 
Structure of human SARS-CoV-2 spike glycoprotein trimer bound by neutralizing antibody C1C-A3 Fab (variable region)
Method: single particle / : Pan J, Abraham J, Shankar S

EMDB-23574: 
Structure of Plasmodium falciparum 20S proteasome with bound bortezomib
Method: single particle / : Liu B, Hanssen E, Leis AP, Xie SC, Morton CJ, Metcalfe RD, Tilley L, Griffin MDW

EMDB-23575: 
Structure of Plasmodium falciparum 20S proteasome with bound MPI-5
Method: single particle / : Hanssen E, Liu B, Leis AP, Xie SC, Morton CJ, Metcalfe RD, Tilley L, Griffin MDW

EMDB-23576: 
Structure of human 20S proteasome with bound MPI-5
Method: single particle / : Hanssen E, Xie SC, Liu B, Leis AP, Morton CJ, Metcalfe RD, Tilley L, Griffin MDW

PDB-7lxt: 
Structure of Plasmodium falciparum 20S proteasome with bound bortezomib
Method: single particle / : Morton CJ, Metcalfe RD, Liu B, Xie SC, Hanssen E, Leis AP, Tilley L, Griffin MDW

PDB-7lxu: 
Structure of Plasmodium falciparum 20S proteasome with bound MPI-5
Method: single particle / : Metcalfe RD, Morton CJ, Xie SC, Liu B, Hanssen E, Leis AP, Tilley L, Griffin MDW

PDB-7lxv: 
Structure of human 20S proteasome with bound MPI-5
Method: single particle / : Metcalfe RD, Morton CJ, Liu B, Xie SC, Hanssen E, Leis AP, Tilley L, Griffin MDW

EMDB-24504: 
Structure of the SARS-CoV-2 S 6P trimer in complex with neutralizing antibody C118 (State 1)
Method: single particle / : Barnes CO, Jette CA, Bjorkman PJ

EMDB-24505: 
Structure of the SARS-CoV-2 S 6P trimer in complex with neutralizing antibody C118 (State 2)
Method: single particle / : Barnes CO, Jette CA, Bjorkman PJ

PDB-7rkv: 
Structure of the SARS-CoV-2 S 6P trimer in complex with neutralizing antibody C118 (State 1)
Method: single particle / : Barnes CO, Jette CA, Bjorkman PJ

EMDB-23312: 
BG505 SOSIP.v5.2 in complex with VRC40.01 and RM19R Fabs
Method: single particle / : Cottrell CA, Torres JL, Wu NR, Ward AB

PDB-7lg6: 
BG505 SOSIP.v5.2 in complex with VRC40.01 and RM19R Fabs
Method: single particle / : Cottrell CA, Torres JL, Wu NR, Ward AB

EMDB-23693: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the human neutralizing antibody Fab fragment, BG10-19
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-23694: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the human neutralizing antibody Fab fragment, BG1-22
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-23695: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the human neutralizing antibody Fab fragment, BG7-15
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-23696: 
Structure of the SARS-CoV-2 S 2P trimer in complex with the human neutralizing antibody Fab fragment, BG7-20
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-23697: 
Structure of the SARS-CoV-2 S 2P trimer in complex with the human neutralizing antibody Fab fragment, BG1-24
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-23424: 
Cryo-EM map of Q23.17_DS-SOSIP in complex with Glycan276-Dependent Broadly Neutralizing Antibody 179NC75 Fab
Method: single particle / : Manne K, Acharya P

PDB-7llk: 
Cryo-EM structure of Q23.17_DS-SOSIP in complex with Glycan276-Dependent Broadly Neutralizing Antibody 179NC75 Fab
Method: single particle / : Manne K, Acharya P

PDB-7m6e: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the human neutralizing antibody Fab fragment, BG10-19
Method: single particle / : Barnes CO, Bjorkman PJ

PDB-7m6f: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the human neutralizing antibody Fab fragment, BG1-22
Method: single particle / : Barnes CO, Bjorkman PJ

PDB-7m6g: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the human neutralizing antibody Fab fragment, BG7-15
Method: single particle / : Barnes CO, Bjorkman PJ

PDB-7m6h: 
Structure of the SARS-CoV-2 S 2P trimer in complex with the human neutralizing antibody Fab fragment, BG7-20
Method: single particle / : Barnes CO, Bjorkman PJ

PDB-7m6i: 
Structure of the SARS-CoV-2 S 2P trimer in complex with the human neutralizing antibody Fab fragment, BG1-24
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-23411: 
Cryo-EM map of BG505 DS-SOSIP in complex with glycan276-dependent broadly neutralizing antibody VRC40.01 Fab
Method: single particle / : Manne K, Acharya P

EMDB-23412: 
Cryo-EM map of BG505 DS-SOSIP in complex with Glycan276-Dependent Broadly Neutralizing Antibody VRC33.01 Fab
Method: single particle / : Manne K, Acharya P

PDB-7ll1: 
Cryo-EM structure of BG505 DS-SOSIP in complex with glycan276-dependent broadly neutralizing antibody VRC40.01 Fab
Method: single particle / : Manne K, Acharya P

PDB-7ll2: 
Cryo-EM structure of BG505 DS-SOSIP in complex with Glycan276-Dependent Broadly Neutralizing Antibody VRC33.01 Fab
Method: single particle / : Manne K, Acharya P

EMDB-21005: 
Negative stain EM map of CODH/ACS in the closed conformation
Method: single particle / : Cohen SE, Brignole EJ, Drennan CL
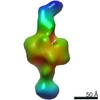
EMDB-21006: 
Negative stain EM map of CODH/ACS in the hyperextended conformation
Method: single particle / : Cohen SE, Brignole EJ, Drennan CL

EMDB-21007: 
Negative stain EM map of CODH/ACS in the half-extended conformation
Method: single particle / : Cohen SE, Brignole EJ, Drennan CL
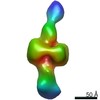
EMDB-21008: 
Negative stain EM map of CODH/ACS in the extended conformation
Method: single particle / : Cohen SE, Brignole EJ, Drennan CL

EMDB-21414: 
Trimeric Complex of Them1 Negative Stain Map
Method: single particle / : Tillman MC, Imai N, Li Y, Khadka M, Okafor CD, Juneja P, Adhiyaman A, Hagen SJ, Cohen DE, Ortlund EA

EMDB-20645: 
MicroED structure of a FIB-milled CypA Crystal
Method: electron crystallography / : Wolff AM, Martynowycz MW, Zhao W, Gonen T, Fraser JS, Thompson MC

PDB-6u5g: 
MicroED structure of a FIB-milled CypA Crystal
Method: electron crystallography / : Wolff AM, Martynowycz MW, Zhao W, Gonen T, Fraser JS, Thompson MC
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
